|
7/25/2023 0 Comments Protein turnover 
The ubiquitin–proteasome system (UPS) is the main pathway for selective protein degradation, which uses a diverse collection of E1, E2, and E3 ubiquitin ligases to add ubiquitin to both cytosolic and nuclear proteins, targeting them for degradation by the proteasome ( 20, 21). Protein degradation occurs via two proteolytic machineries: the lysosome and the proteasome. Molecular chaperones, insertases, and translocases control maturation of nascent polypeptide chains, and post-translational modifications are added to proteins in the secretory pathway or through signaling cascades ( 17, 18, 19). mRNA translation is controlled by regulatory motifs in mRNA nucleotide sequences these sequences are bound by RNA-binding proteins and small RNA guides (such as microRNAs) to modulate their expression ( 12, 13, 14, 15, 16). Although protein production includes all of the processes that precede mRNA translation, including RNA transcription, maturation, and processing, in this review, we will focus on the time frame between protein synthesis and degradation.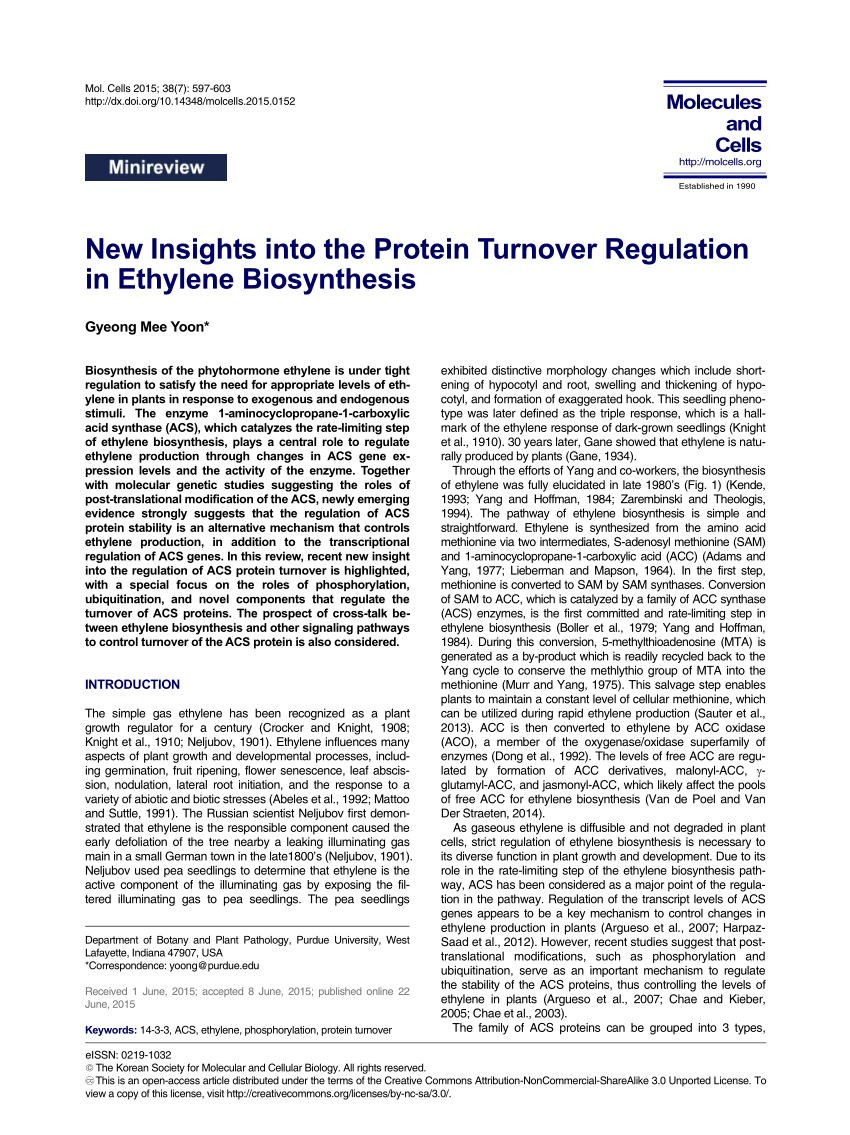 
Protein turnover is monitored and regulated by several cellular surveillance systems. Probing the kinetics of proteome-wide protein turnover lends insight into how cells perform crucial functions such as differentiation and stress response in both normal and disease contexts, and can illuminate the guiding principles that underlie the regulation of protein turnover across protein families, cell types, and species. Accordingly, disruption of proteostasis contributes to the pathophysiology of a variety of disease states, most notably neurodegenerative disorders and cancer ( 11). 
These processes also ensure that misfolded, aged, or damaged proteins are removed from the cellular protein pool as needed ( 11). Proteostatic mechanisms are some of the cell’s most essential processes, as they ensure that functional proteins are maintained at their correct concentrations and in the proper locations needed for cellular activities to proceed ( 8, 9, 10). Protein turnover is regulated by several tightly controlled processes that help facilitate protein homoeostasis, also known as proteostasis ( 1, 2, 3, 4, 5, 6, 7). In all cells, proteins are continuously produced and degraded, a process referred to as protein turnover. We highlight important considerations for designing proteome turnover experiments, key biological findings regarding the conserved principles of proteome turnover regulation, and future perspectives for both technological and biological investigation. Here, we outline historical and current approaches to measuring the kinetics of protein turnover on a proteome-wide scale in both steady-state and dynamic systems, with an emphasis on metabolic tracing using stable isotope–labeled amino acids. Thus, probing the kinetics and dynamics of protein turnover lends insight into how cells regulate essential processes such as growth, differentiation, and stress response. At a steady state, protein turnover is constant to maintain protein homeostasis, but in dynamic responses, proteins change their rates of synthesis and degradation to adjust their proteomes to internal or external stimuli. Collectively, the processes of protein synthesis and degradation are referred to as protein turnover. In all cells, proteins are continuously synthesized and degraded to maintain protein homeostasis and modify gene expression levels in response to stimuli.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
